| Ding et al. report on the use of deoxyribonucleic acid (DNA) origami as a scaffold for the assembly of gold nanoparticles into a carefully organized structure to create, ideally, a device with a strongly enhanced plasmonic resonance. |
Reviewed by J. Alexander Liddle, National Institute of Standards and Technology
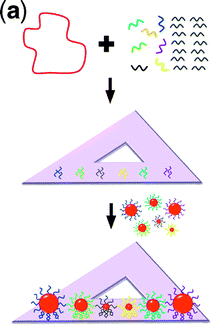
Ding et al. chose to realize a plasmonic nanolens structure first analyzed by Li et al. (Physical Review Letters (2003)). Those authors described how a chain of three self-similar gold nanoparticles can give rise to field enhancements of up to a thousand-fold, depending on the wavelength of light used and the precise arrangement of the nanoparticles. The greatest field enhancements are obtained when the smallest particle is on the order of 10 nm in diameter with particle-to-particle spacings as small as 9 nm. These dimensions are difficult to obtain reliably with lithographic methods, making an alternative approach to fabrication attractive.

Once they had produced origami with the plasmonic devices, the authors conducted preliminary measurements of their optical response, observing a 5 nm shift in the location of the plasmon-band peak in the ultraviolet-visible absorbance spectrum, showing some level of plasmonic interaction. One question that always arises in the context of trying to model and measure the plasmonic response of a nanostructure is: how sensitive is the response to the precise details of particle size, shape and spacing? This is particularly important given the difficulty of controlling the morphology of such small objects. Another key question is: how will the completed nanoparticle structures be integrated into larger-scale devices? Fortunately, this is an active area of research with recent papers by Kershner et al. and Hung et al. (Nature Nanotechnology (2009)) focusing on the use of lithographically-patterned, chemically-functionalized surfaces to control the placement and orientation of origami. Finally, and perhaps most critical, as far as applications directed towards integration into silicon electronics, is the question of what are the fundamental limits in terms of the overall yields that can be achieved?
Images reproduced with permission from Ding B, et al. 2010. Gold Nanoparticle Self-Similar Chain Structure Organized by DNA Origami. J. Am. Chem. Soc. 132 (10): 3248–3249. DOI:10.1021/ja910119. Copyright 2010 American Chemical Society.
This work is licensed under a Creative Commons Attribution-NonCommercial 3.0 Unported.
